Building biomedical ontologies for a Life Sciences Semantic Web
3S Biology Summer School (Trento, 2014)
Mikel Egaña Aranguren
http://mikeleganaaranguren.com / mikel.egana@ehu.es

http://mikeleganaaranguren.wordpress.com/teaching/
http://mikel-egana-aranguren.github.io/3SBiologyTrento2014/
http://github.com/mikel-egana-aranguren/3SBiologyTrento2014

Life Sciences Semantic Web
Life Sciences Semantic Web
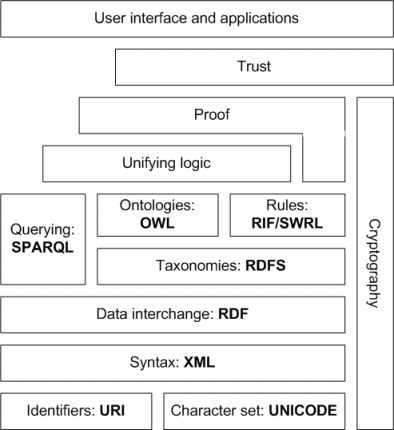
Semantic Web Stack
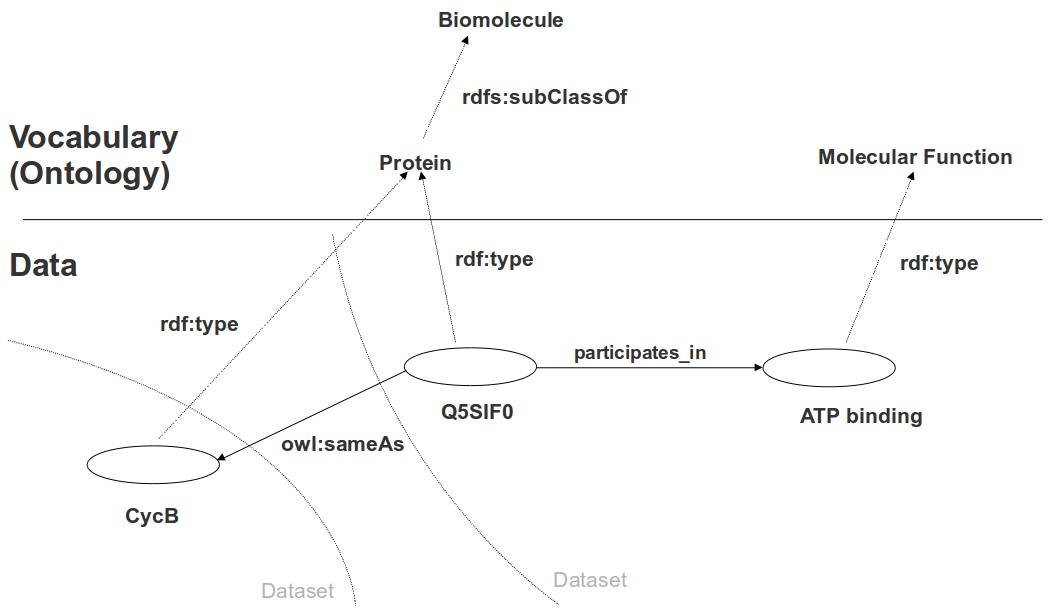
Ontologies and Semantic Web
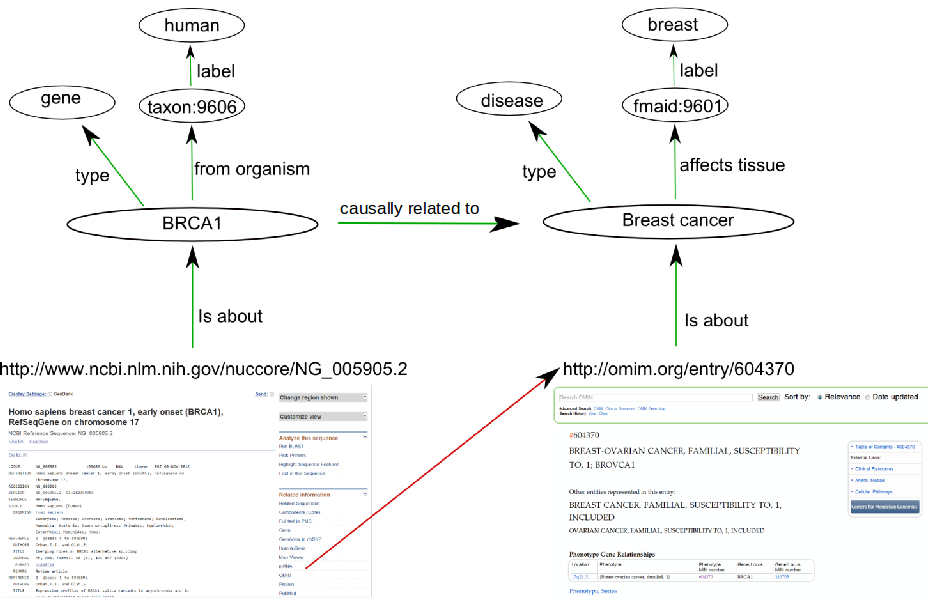
Biomedical Ontologies
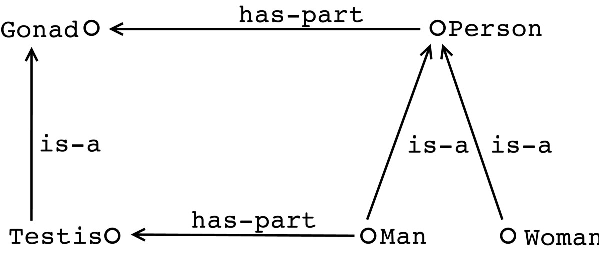
Functions
"Schema" for Knowledge Base
Vocabulary for publishing Linked Data
Vocabulary for common database annotation (e.g. GO)
Text mining
Web Services description (e.g. SADI)
Data analysis
...
Gene Ontology
Controlled vocabulary for describing molecular function, cellular component and biological process of gene products
Database integration through Gene Association Files (GAF), data analysis (e.g. Term Enrichment), ...
Ontology engineering languages
OBO format ("DEPRECATED"!)

Web Ontology Language (OWL)
A W3C official standard for creating ontologies in a Semantic Web setting with computationally precise and formal semantics
OWL
OWL is based in Description Logics (DL)
Automated reasoning can be used for inference: "if a cow is a kind of animal and animal is a kind of organism, then cow is a kind of organism"
OWL syntax
For computers: RDF/XML, OWL/XML, ...

For human beings: Manchester OWL Syntax, functional, ...

OWL semantics
An OWL ontology is comprissed of:
- Entities: the elements of the knowledge domain, identified with URIs, added by the ontology curator ("protein", "participates_in", ...)
- Axioms: they relate entities through the logical standard vocabulary offered by OWL, creating asssertions about the knwoledge domain ("In order to be a protein an entity must participate in at least one metabolic process")
An ontology can import (owl:import) other ontology and refer to its entities through axioms
OWL entities
- Individuals
- Clases
- Properties
- Object
- Anotation
- Data
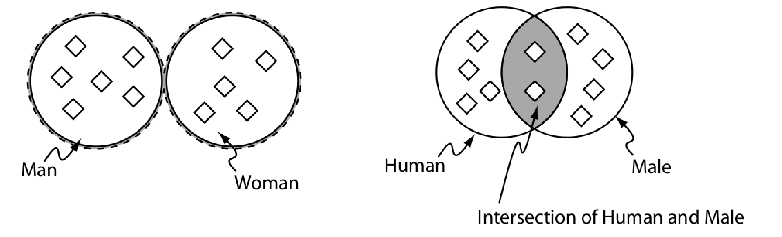
Individuals
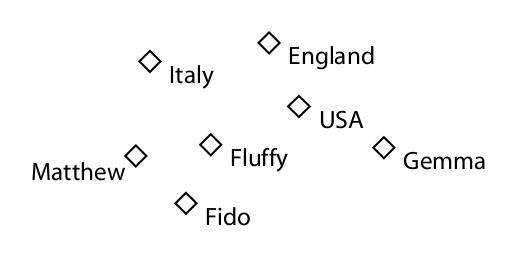
Properties
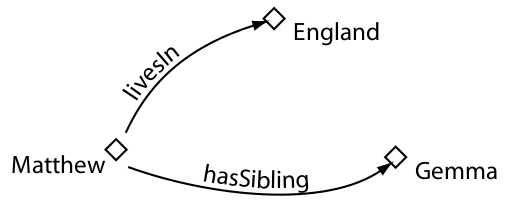
Classes
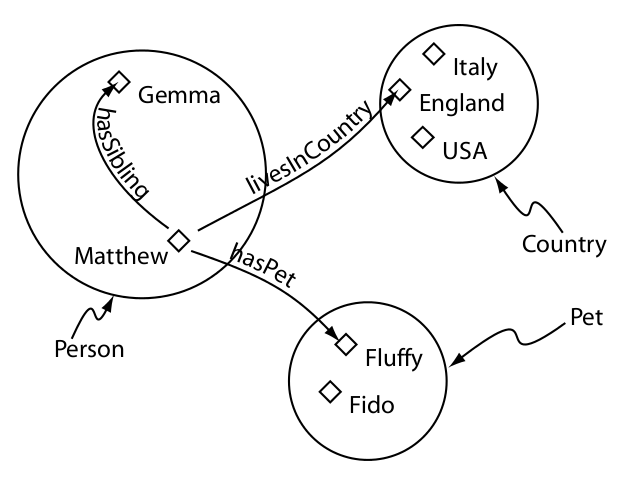
Classes
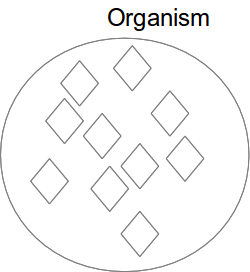
Class subclass (subsumption)
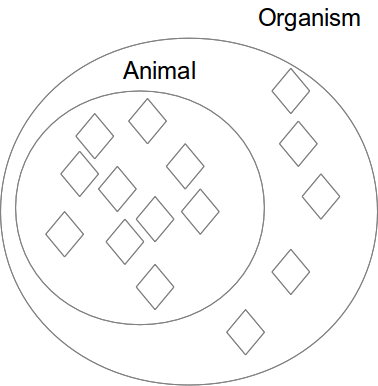
Equivalent classes
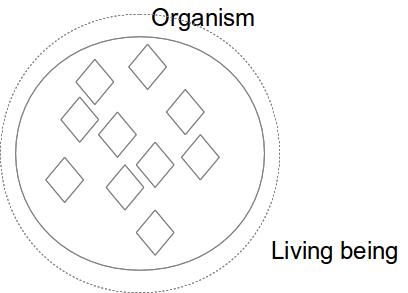
Class hierarchy (Taxonomy)
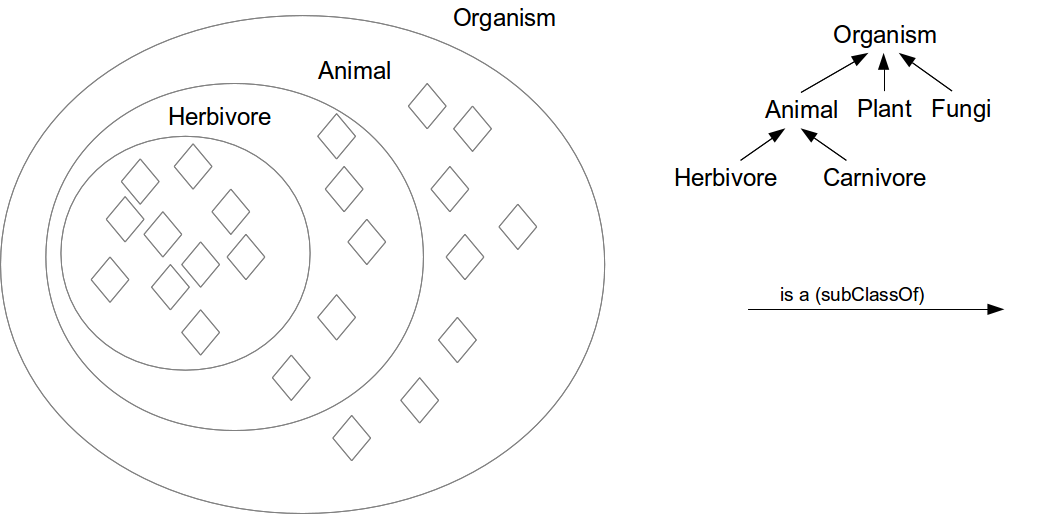
Necessary conditions
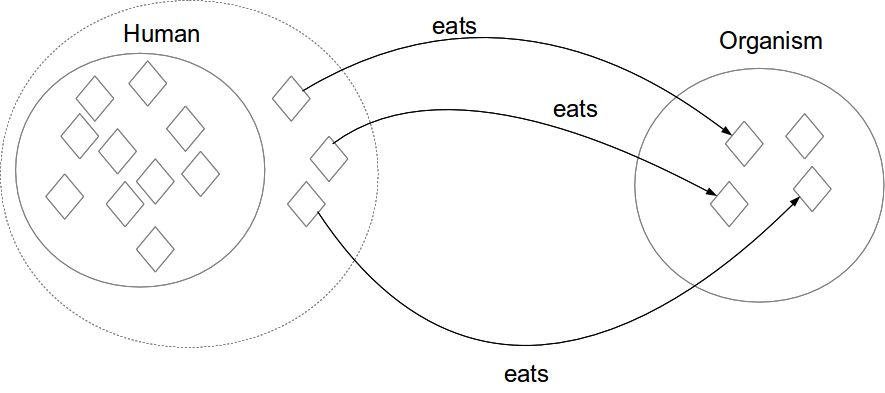
Necessary and sufficient conditions
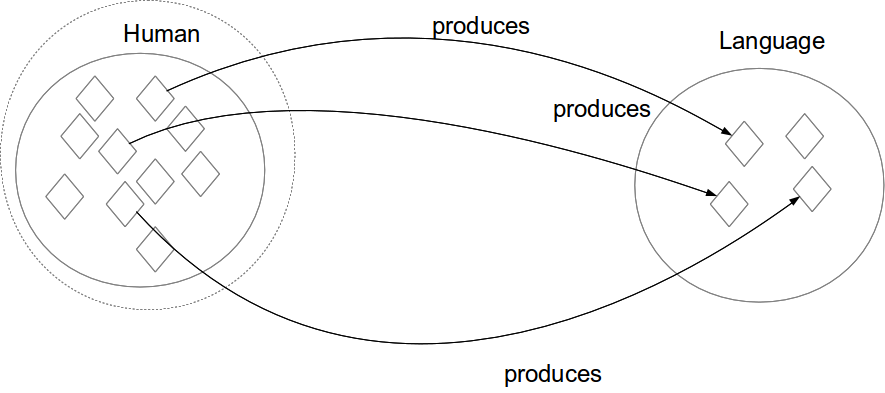
Existential restriction (some)
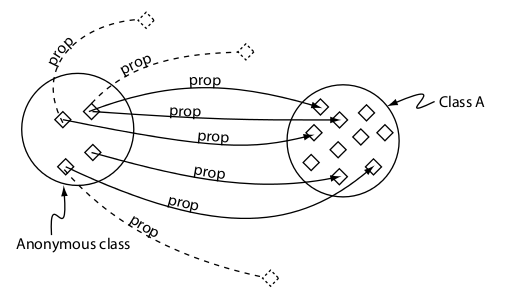
Universal restriction (only)
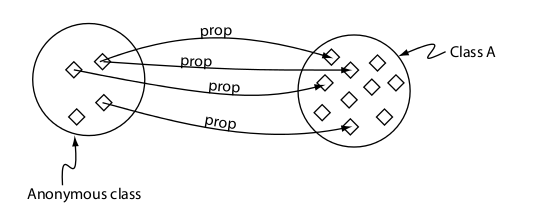
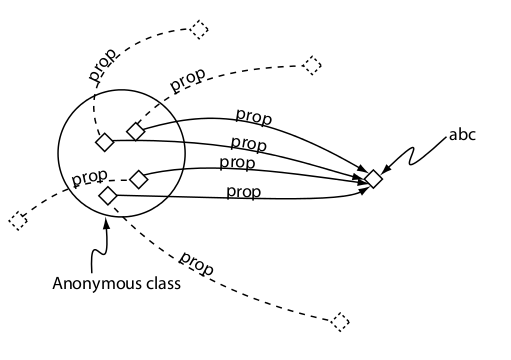
Restriction to a individual (value)
Cardinality constraints

(+ QCR!) Manchester tutorial
More axioms for classes
disjointFrom
booleans: not, or, and
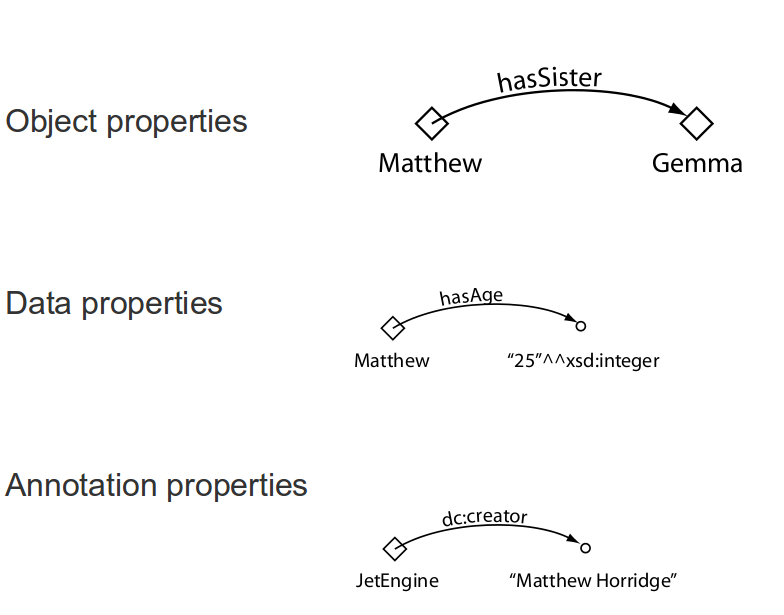
OWL properties
Object property features
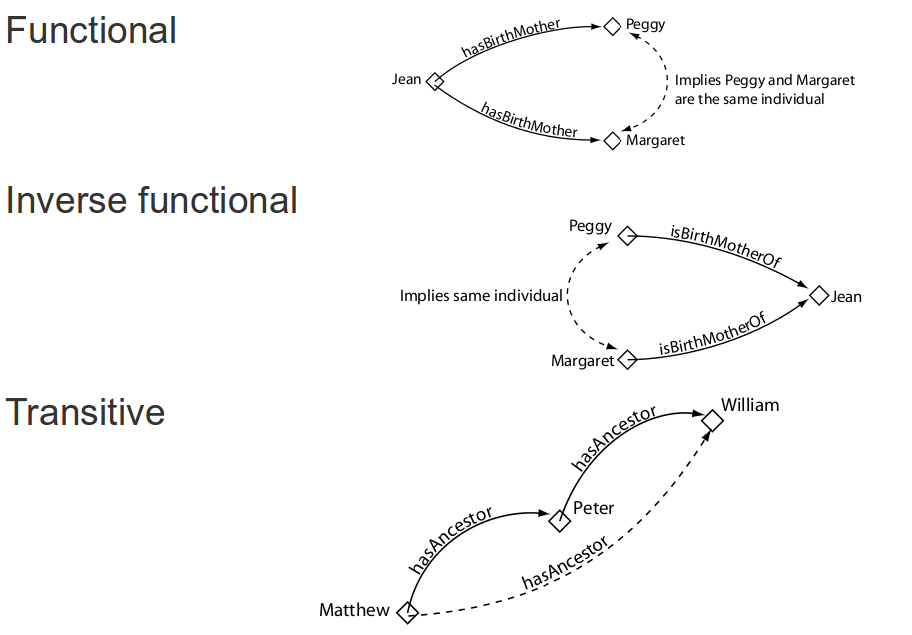
Object property features
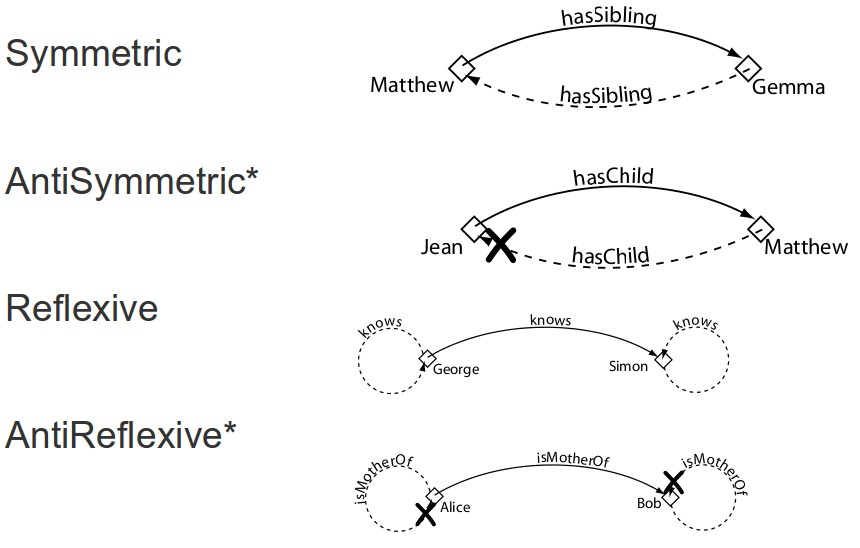
Object property features
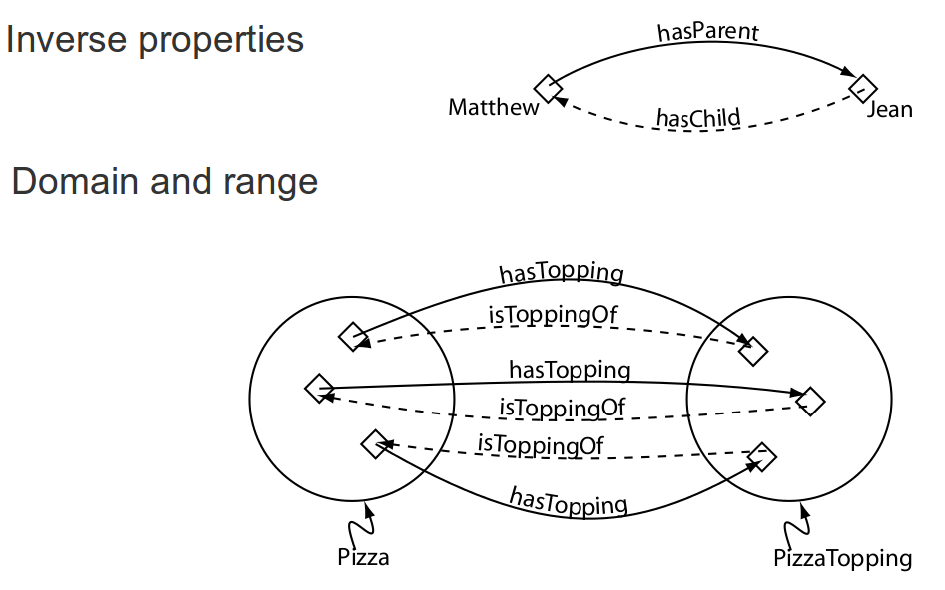
Annotation properties
Natural language information that can be attached to entities (properties, individuals, classes), axioms, ontologies
No semantics
rdfs:label, rdfs:comment, dublin core, create new ones ...
Individuals
Member of one or more classes (Types)
SameAs or DifferentFrom other individuals
Binary relations with other individuals or data (triples), positive or negative
Complex expressions
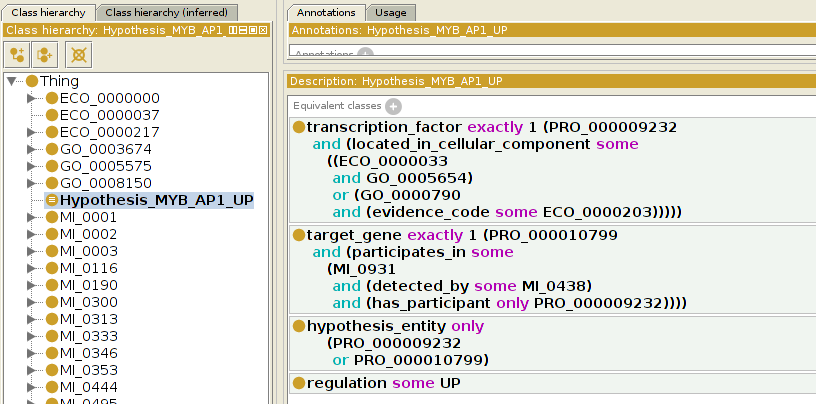
Automated reasoning
A reasoner will infer the "new" (*?!) axioms that the axioms we have stated in the ontology imply
The reasoner will infer all the implied axioms: it is useful to deal with complex knowledge
Open World Assumption (OWA)
(Lack of) Unique Name Assumption (and use of owl:sameAs)
Reasoning common tasks
Maintain a taxonomy
Check consistency
Clasify entities and pose queries against the ontology
Maintain a taxonomy
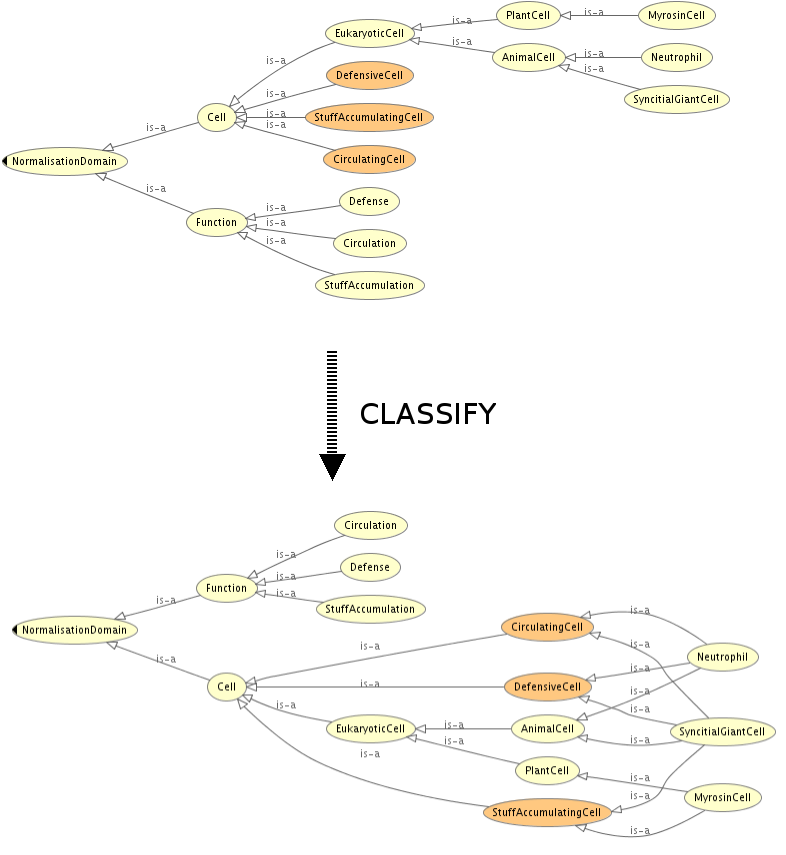
Consistency
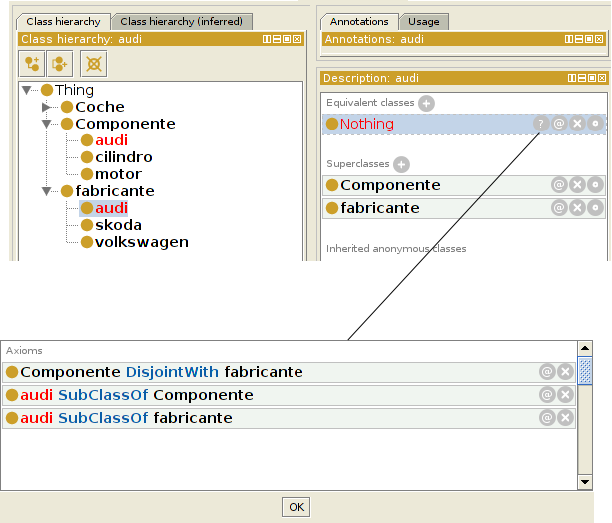
Clasify entities, queries
Clasify entities: given an entity (e.g. an individual with certain relations), infer how it relates to other entities (e.g. to which classes it belongs) and the consistency of the model including the new entity
A query is an anonymous class that we classify against the ontology as if it were and entity
Ontology engineering tools
APIs
...
Ontology editors
...
Automated Reasoners
...
Ontology engineering practices
Scope
Define a scope: domain (e.g. Cell Cycle), application (e.g. BioPAX), ...
Stick to it
Identifiers
Define a Semantic Web friendly URI scheme:
Cool URIs for the Semantic Web
Linked Data URIs
...
Stick to it
Identifiers
Use common identifiers (e.g. identifiers.org)
Use alphanumeric identifiers in URIs and rdfs:label for common names
Use permament URLs (e.g. http://purl.oclc.org)
Interoperability
Commonly used Upper Level Ontology (e.g. BFO)
Or your own one, but be explicit
Commonly used properties (e.g. OBO Relations Ontology)
Or your own ones, but be explicit
Interoperability
Reuse existing ontologies, extend them if needed ...
... and only then, and if completely necessary, create new entities
Link to other ontologies with owl:equivalentClass, rdf:type, rdf:subClassOf, ...
Automate
Ontology manipulation (create entities, add/remove axioms):
Releases: OORT (OWLtools)
Automate
Use a VCS as usual (e.g. GitHub), but be aware of perils of managing OWL in a VCS
Apply automated reasoning after every change
Document
Use annotation properties everywhere a lot:
rdfs:label
rdfs:comment
Dublin Core
your own ones
Modelling
The more explicit (axioms), the better (within reason). Reasoner always knows better than you
Follow Ontology Design Patterns (ODPs) and create/share your own ones: odps.sf.net, ontologydesignpatterns.org
Pitfalls: OntOlogy Pitfall Scanner (OOPS!), catalogue of ontology pitfalls
Community
OBO Foundry: Open, Common shared syntax, Unique identifier space, Versions, Delineated content, Definitions, OBO Relation Ontology, Well documented, Users, Collaboratively
An actual (not so good :P) example
BIOMO (Biological Observation Matrix Ontology):